Jug Group
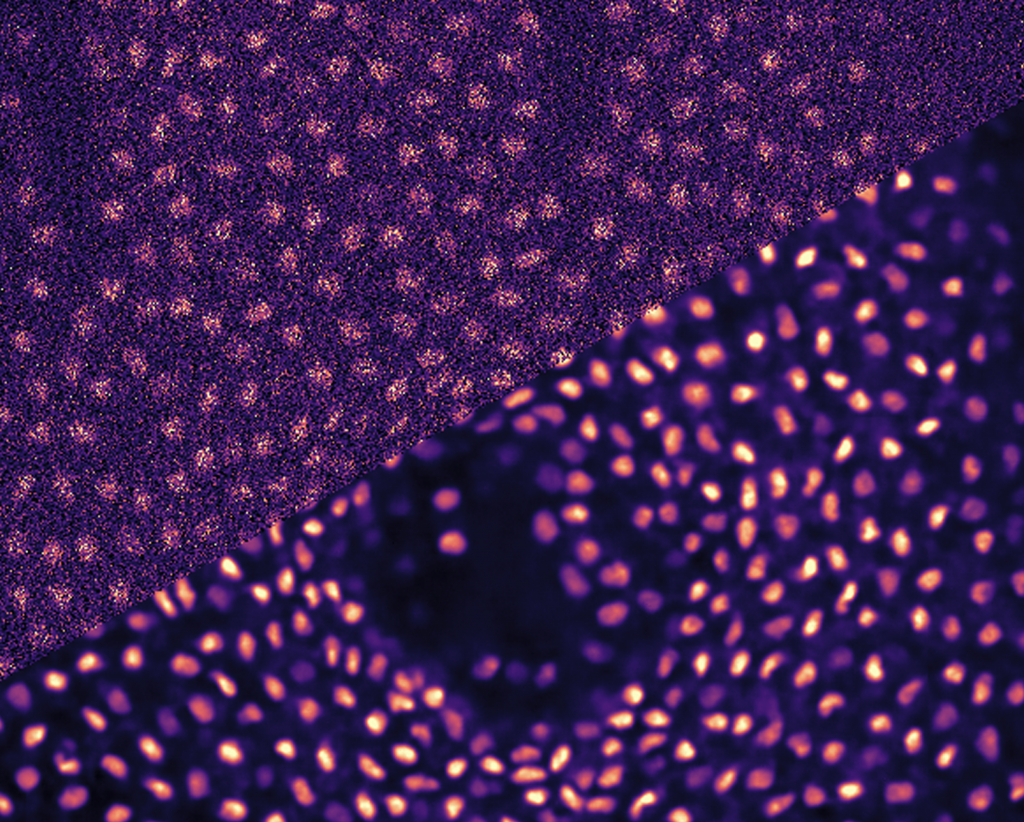
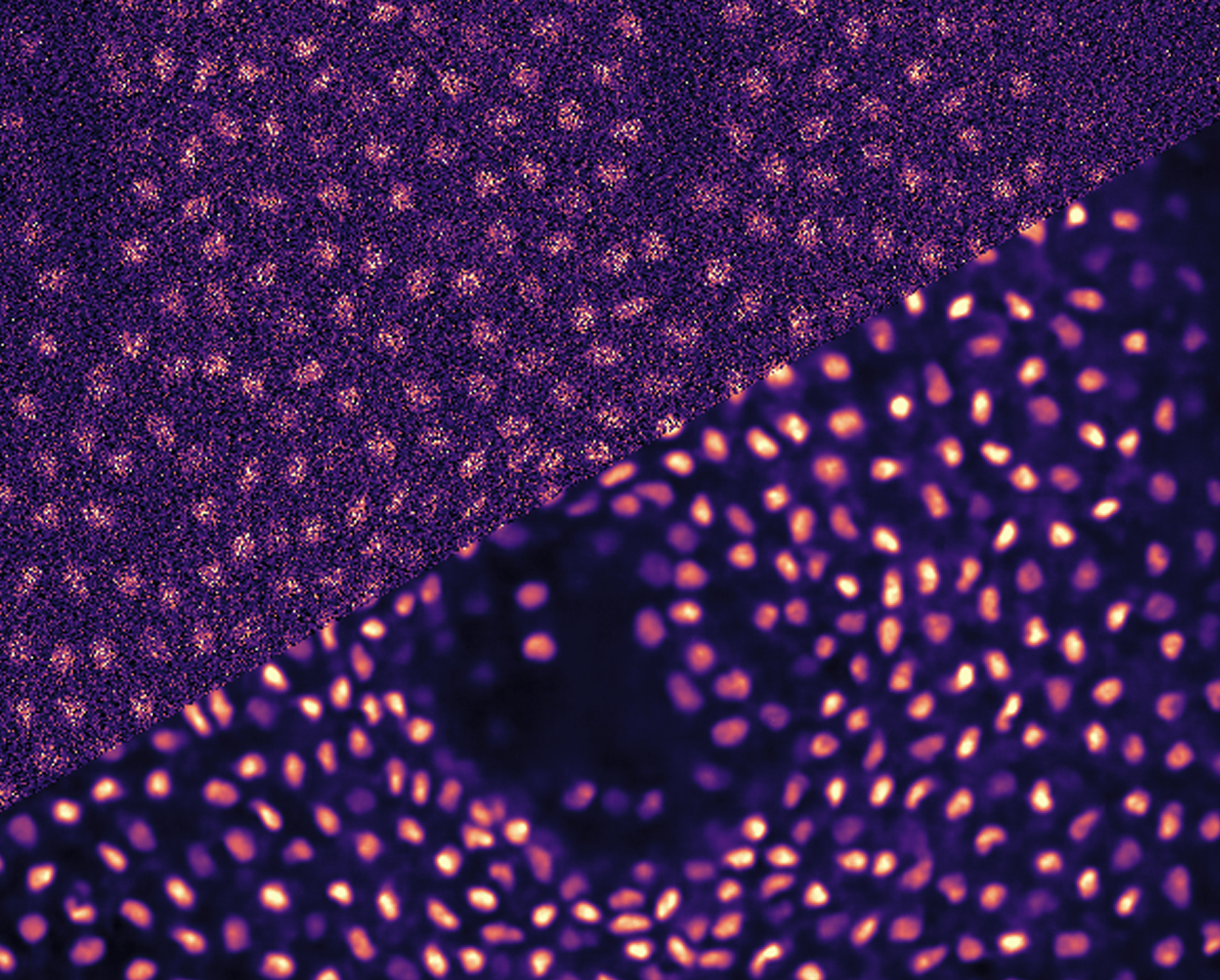
The research conducted in the Jug Group is pushing the boundary of what image analyses and machine learning can do for quantifying biological (image) data. The common denominator of such projects is the indisputable necessity to analyse large amounts of light microscopy data without causing impossible amounts of manual data curation and data processing to life-science researchers (aka our users and collaborators).
From a computational point of view we are interested in (i) image denoising and restoration, (ii) object segmentation and tracking, and (iii) analysis modules for tasks such as object detection, nD image registration/ deformation, etc.
Next to developing novel machine learned and algorithmic solutions, the Jug Group also has a strong emphasis on casting these methods into modular and reusable software packages. This work happens to a large degree in the context of Fiji and the trending scientific python world. We strongly believe that the power and flexibility of the image.sc community and their very rich universe of open source solutions is of a value and utility to the life-science community at large that can hardly be underestimated.
Group members
-
 Florian Jug
Florian Jug
Research Group Leader -
 Ashesh Ashesh
Ashesh Ashesh
PhD Student -
 Federico Carrara
Federico Carrara
PhD Student -
 Vera Galinova
Vera Galinova
Bioimage Analyst and Research Software Engineer -
 Edoardo Giacomello
Edoardo Giacomello
Machine Learning Specialist -
 Aman Dhanraj Kukde
Aman Dhanraj Kukde
Postgraduate Fellow -
 Sheida Rahnamai Kordasiabi
Sheida Rahnamai Kordasiabi
PhD Student -
 Anirban Ray
Anirban Ray
PhD Student -
 Ruggiero Santeramo
Ruggiero Santeramo
Machine Learning Specialist -
 Mehdi Seifi
Mehdi Seifi
Bioimage Analyst and Research Software Engineer -
 Diya Srivastava
Diya Srivastava
Data Scientist and Research Software Engineer -
 Igor Zubarev
Igor Zubarev
Bioimage Analyst and Research Software Engineer
Publications
-
06/2020 - arXiv
DivNoising: Diversity Denoising with Fully Convolutional Variational Autoencoders
Deep Learning based methods have emerged as the indisputable leaders for virtually all image restoration tasks. Especially in the domain of microscopy images, various content-aware image restoration (CARE) approaches are now used to improve the interpretability of acquired data. But there are limitations to what can be restored in corrupted images, and any given method […]
-
06/2020 - arXiv
DenoiSeg: Joint Denoising and Segmentation
Microscopy image analysis often requires the segmentation of objects, but training data for this task is typically scarce and hard to obtain. Here we propose DenoiSeg, a new method that can be trained end-to-end on only a few annotated ground truth segmentations. We achieve this by extending Noise2Void, a self-supervised denoising scheme that can be […]