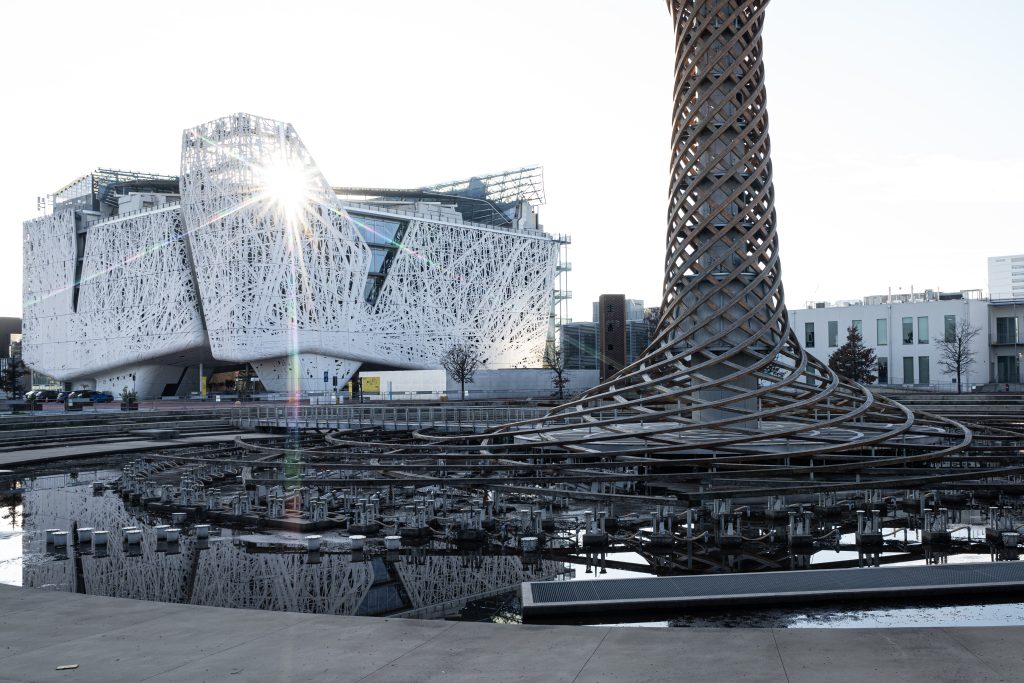
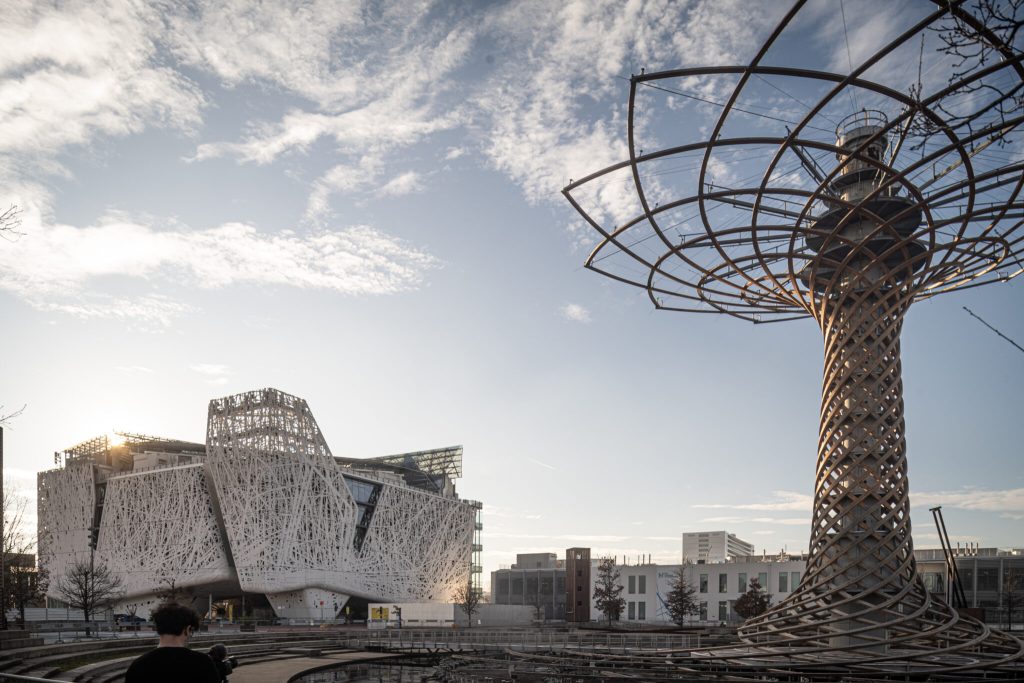
Preserving and consolidating biomedical databases – Human Technopole signs MoU with the Department of Biology of the University of Rome “Tor Vergata”

Milan, 1 April – Human Technopole and the Department of Biology of the University of Rome “Tor Vergata” have signed an MoU that demonstrates their joint commitment towards the continued development of two databases, MINT and SIGNOR, which over time have become fundamental resources for the European bioinformatics community.
Around 20 years ago the Molecular Genetics Group in Tor Vergata, led by Prof. Cesareni, initiated the development of two data resources, the Molecular INTeraction (MINT) and, more recently, the SIGnalling Network Open Resource (SIGNOR) databases, aimed at capturing literature information about physical and functional molecular interactions respectively.
Discussions between Human Technopole and the Department of Biology of “Tor Vergata” have been ongoing since 2019 in an effort to ensure the continuity of the two databases, which require the work of expert staff who mine and annotate information from scientific literature. Scientists from all over the world can access the databases in the spirit of openness that characterises scientific research.
Over time, MINT has become increasingly important within the scientific community and is now part of the ELIXIR network, the intergovernmental organisation that brings together life science resources from across Europe. It is currently the only Italian data resource to be included among the ELIXIR Core Data Resources.
The cooperation between the two organisations towards the continued development of the MINT and SIGNOR datasets is envisioned to take different shapes, and to be pursued through the work of dedicated informaticians and curators. A further agreement will regulate specific aspects of the collaboration, including the future governance structure and policy on the use of the MINT and SIGNOR databases to benefit the wider research community and thus facilitating new breakthroughs in biomedicine.
“The close interaction between the databases developed by the Department of Biology in Tor Vergata and the Human Technopole projects will ensure the continuity of these important biological resources while at the same time contributing to advance the Human Technopole mission of linking genetic data to disease phenotypes” comments Prof. Cesareni.
The data sets. MINT is an established resource that is a founding member of the international iMEX consortium of federated databases. The iMEX consortium aims at coordinating the capture – in a structured and easily accessible format – of information on experimentally verified protein-protein interactions. Whereas the more recent SIGNOR, using a similar approach, specifically focusses on causal relationships (i.e. functional interactions) between proteins and other biological entities involved in signal transduction. The MoU also involves collaborative work on two ancillary disease-focussed resources, DISNOR (for genetic diseases) and CancerGeneNet (for cancers) that have been developed to support genetic data interpretation and personalised medicine. DISNOR in particular combines the causal interaction information annotated in SIGNOR with protein interaction data to generate and explore disease pathways. CancerGeneNet aims at connecting genes that are frequently mutated in cancers to cancer phenotypes, thereby linking driver genes to cancer hallmarks.