Genomics Facility welcomes first sequencers
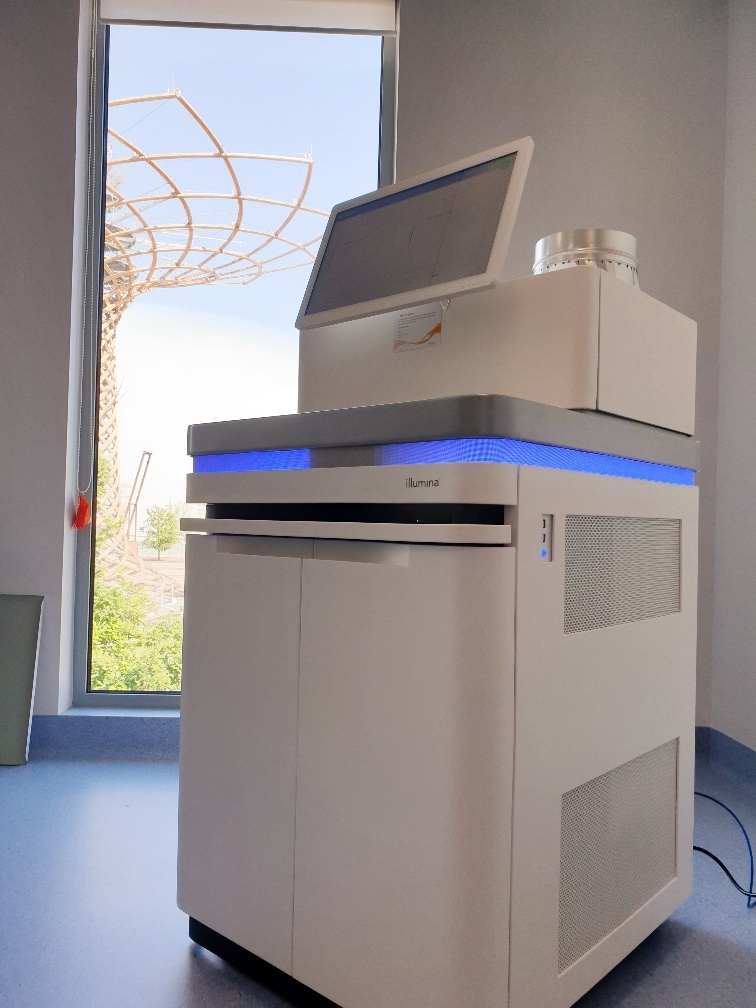

Last week the first Next Generation sequencers of Human Technopole’s Genomics Facility were delivered and installed on the first floor of Incubator Lab 1.
Clelia Peano is the newly appointed Senior Manager of High-Throughput Sequencing Operations and we asked her some questions on this new, exciting development on campus.
What is the aim of HT’s Genomics Facility?
The Genomics Facility is one of Human Technopole’s six core facilities, a strategic infrastructure for the implementation of our mission and projects. It will work alongside all of HT’s research centres providing high throughput analysis potential for large projects and single-cell level analysis, allowing to analyse various types of samples with an increasing level of sensitivity and specificity.
Once fully operational, the Facility will be able to provide state-of-the-art and innovative services in Genomics, Transcriptomics, Epigenomics, and Metagenomics fields of research.
What is Next Generation sequencing and how does it work?
Next-generation sequencing (NGS) is a massively parallel sequencing technology that is used to determine the order of nucleotides in entire genomes or targeted regions of DNA or RNA. It has revolutionized the biological sciences, allowing labs to perform a wide variety of applications and study biological systems at a level never before possible.
Next-generation sequencing generates masses of DNA sequencing data and is both less expensive and less time-consuming than traditional Sanger sequencing. For example, certain sequencing systems can deliver data output ranging from 300 kilobases up to multiple terabases in a single run, depending on instrument type and configuration.
Next Generation Sequencing technology has fundamentally changed the kinds of questions scientists can ask and answer. Innovative sample preparation and data analysis options enable a broad range of applications. For example, NGS allows labs to:
- rapidly sequence whole genomes;
- use RNA sequencing (RNA-Seq) to discover novel RNA variants and splice sites, or quantify mRNAs for gene expression analysis;
- analyse epigenetic factors such as genome-wide DNA methylation and DNA-protein interactions;
- deeply sequence target regions;
- sequence cancer samples to study rare somatic variants, tumor subclones, and more;
- identify novel pathogens;
- study the human microbiome.
The workflows includes three basic steps:
1. Library Preparation: The sequencing library is prepared by random fragmentation of the DNA or cDNA sample, followed by 5′ and 3′ adapter ligation. Alternatively, “tagmentation” combines the fragmentation and ligation reactions into a single step that greatly increases the efficiency of the library preparation process. Adapter-ligated fragments are then PCR amplified and purified.
2. Cluster Generation: For cluster generation, the library is loaded into a flow cell where fragments are captured on a lawn of surface-bound oligos complementary to the library adapters. Each fragment is then amplified into distinct, clonal clusters through bridge amplification. When cluster generation is complete, the templates are ready for sequencing.
3. Sequencing: NGS sequencing utilizes a fundamentally different approach from the classic Sanger chain-termination method. It leverages sequencing by synthesis (SBS) technology – tracking the addition of labelled nucleotides as the DNA chain is copied – in a massively parallel fashion. Nucleotides tagged with fluorescent dyes and a cleavable blocking group are washed over the flow cell; if they are incorporated, the cluster fluoresces with the wavelength associated with that particular nucleotide and the fluorescent signal is acquired and registered by the sequencer. The dye and blocking group are then cleaved to allow the incorporation of the next nucleotide. As all four reversible terminator–bound dNTPs are present during each sequencing cycle, natural competition minimizes incorporation bias and greatly reduces raw error rates compared to other technologies.
What is your role within the Facility?
As Senior Manager of High-Throughput Sequencing Operations, I am in charge of setting up and implementing all the high throughput sequencing services and applications within a two-year frame, thus making the Genomics Facility team and platforms fully operative in all the Genomics investigation areas.
I will coordinate the Genomics Facility team and interact with researchers of the Genomics Research Centre and of the Neurogenomics Research Centre to provide support in planning and realising high throughput sequencing experiments. I will be responsible of all the labs and instruments of the Genomics Facility and of implementing the most innovative sequencing technologies in the facility workflow.
You joined HT very recently – welcome! How is it going so far?
My first month in HT was very exciting, I had the opportunity to meet many new colleagues and to start working to the Genomics Facility set up from the very beginning with new instruments installations and Team building.
My impression up to date is really positive, people are enthusiastic and very kind in general and it is clear that everyone is working in the same direction, sharing the same vision. I am confident that everything is going to be all right and that soon we will start making good science all together.