Nucleolar organisation drives ribosome biogenesis
Ribosomes are complex molecular machines that produce proteins, the key building blocks of all cells. They consist of a small and a large subunit that mature separately and subsequently come together to form a functional ribosome. Ribosome biogenesis is complex: it starts with the synthesis of small- and large-subunit precursors in the nucleolus – a defined sub compartment of the cell nucleus. Nucleoli are very peculiar organelles that are formed through condensation of proteins and other ribosome biogenesis factors into liquid droplets, a process called liquid-liquid phase separation. These liquid-like condensates are membrane-less, yet they have an internal structure, such as the granular component, where immature ribosome assembly is thought to occur.
Defective ribosome biogenesis has been linked to several human diseases. Therefore, understanding the early steps of ribosome formation would be instrumental to elucidating their molecular basis. Although several studies have identified snapshots of nuclear ribosome biogenesis through the structural analysis of isolated ribosomal precursors, it remains unclear how pre-ribosomes are assembled in their native environment.
Philipp Erdmann, Group Leader at the HT Centre for Structural Biology, and his co-workers have taken advantage of an advanced microscopy technique called Cryo-Electron Microscopy (cryo-EM) to analyse the structure of intact nucleoli from the green alga Chlamydomonas reinhardtii and address this fundamental issue. Chlamydomonas is a simple single-cell organism that nonetheless has many ribosome biogenesis factors in common with mammalian cells. The authors have shown that nucleoli, despite their liquid-like nature and their lack of membranes, have an architecture that is important for ribosome biogenesis. The results of their research are now published in Nature Communications.
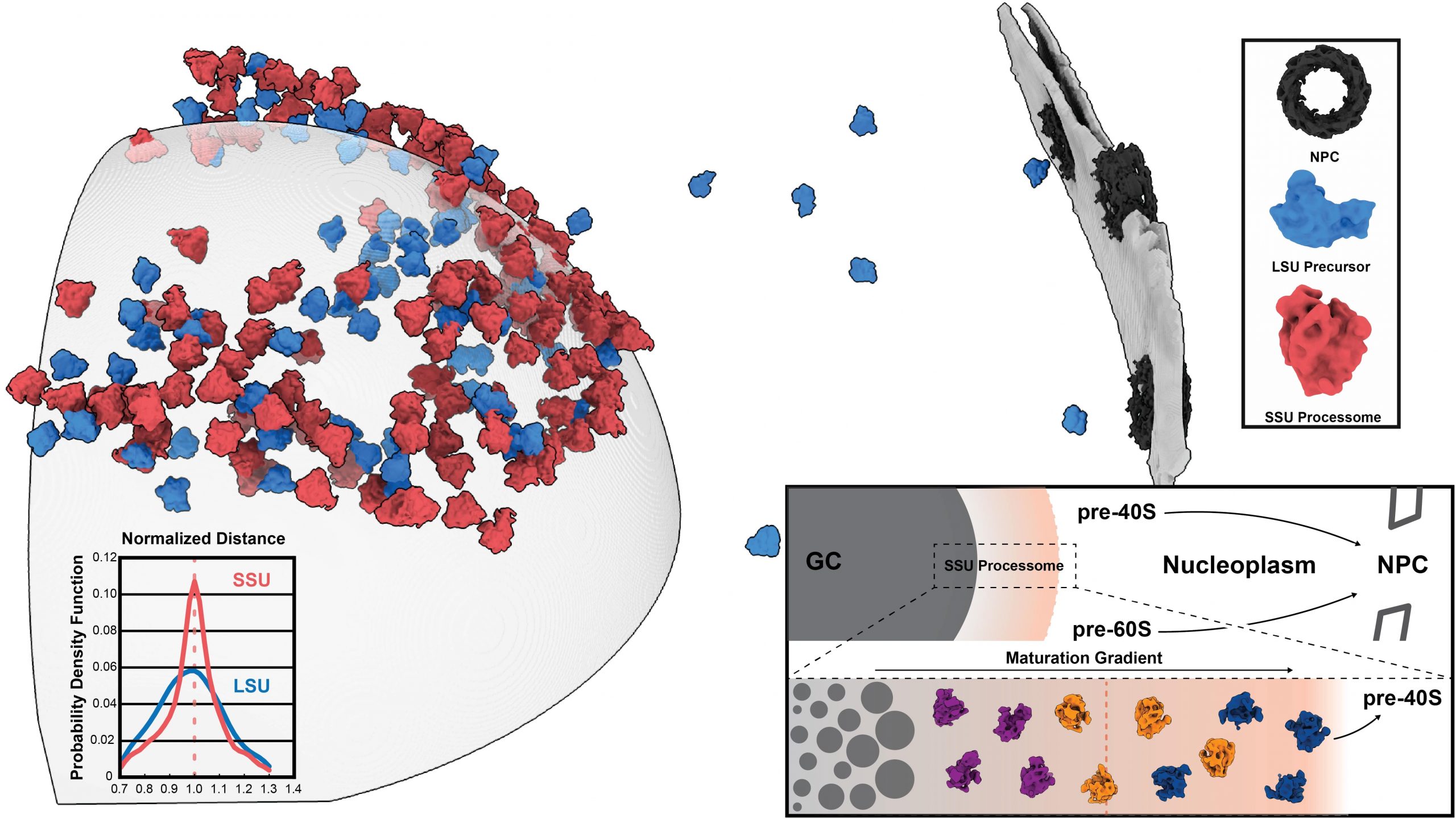
Erdmann et al. rapidly cooled Chlamydomonas cells to very low temperatures and collected thousands of images of frozen nucleoli at various angles to reconstruct their 3D structure, a process called cryo-electron tomography (Cryo-ET). Cryo-ET allowed them to image nucleoli in their native environment in 3D, circumventing artifacts due to nucleoli isolation. Unexpectedly, they found that the small and large ribosomal precursor particles were differentially distributed inside and outside the nucleolus.
By further investigating the architecture of Chlamydomonas nucleoli, the researchers found that mass and size of pre-ribosomal particles increased progressively when moving from the centre to the surface of the nucleolus. They therefore hypothesized that this small-to-large particle gradient could be functionally linked to the maturation of ribosomal precursors. Consistent with this hypothesis, Erdmann and colleagues identified three major classes for both the small- and large-subunit precursors at the nucleolus surface. Comparison of the 3D reconstructions with previously published structures revealed that the intermediates found close to the nucleolar surface resembled more mature states, thus arguing that the observed gradient of pre-ribosomal particles provides a spatiotemporal snapshot of ribosomal precursor maturation.
In summary, Erdmann et al. propose a novel model of ribosome biogenesis whereby pre-ribosomal subunits mature progressively as they move through the surface of the nucleolus. Importantly, the authors show that the precursors of the small ribosomal subunit are those defining the transition of ribosomal particles from the liquid-like nucleolar environment to the nucleoplasm, a key step for ribosome maturation. While further studies will be required to understand how the granular component controls the localisation of other precursors at the nucleolar surface, this study provides the first molecular- resolution evidence that spatial localisation and ribosome maturation are intertwined. These findings will also pave the way to understand how the transition from nucleolar liquid-like state to the cytoplasmic environment affects ribosome biogenesis.